Welcome to Space Race, where we explore the impact of breakthroughs in science research on the intensifying global contest of superpowers in the 21st century. Subscribe for weekly breakdowns of the scientific breakthroughs that will shape the future.
In this week’s newsletter we discuss a creeping issue which threatens to derail the amazing scientific progress underway today — the crisis of reproducibility. We’ll explore why this crisis exists, and what we could do about it. This post is inspired by a conversation I had with angel investor and one-time Stanford PhD student Balaji Srinivasan.
Most of this blog is about the incredible scientific progress going on today, but there is a problem that puts all these advances on shaky footing: many of the results published today will never be independently reproduced by other researchers. Reproducibility is a fundamental pillar of the scientific method, and by making this choice, the scientific community (and academia broadly) is choosing a dangerous path — but it doesn’t need to be this way.
Let’s begin by assessing the ecosystem in which scientists operate to see what incentives drive them. It is tempting to imagine that scientific breakthroughs happen in isolation. We often picture a lone genius toiling away in obscurity until the eureka moment. While that may have been true centuries ago — think of polymaths such as Al-Khwarizmi, Galileo, and Newton — the story today is different. Science has advanced to a level of sophistication where it is impossible for individual geniuses to have any meaningful impact. Galileo could toil on perfecting telescopes in his workshop, but it takes the full resources of a corporate titan like Google to build even a toy quantum computer. For today’s breakthroughs, we need not only geniuses, but a full research ecosystem to support them.
There are two minimum requirements for an effective research ecosystem:
It needs have a way to identify promising areas of research.
It needs to be able to supply the necessary resources (funding, specialised equipment etc.) to make breakthroughs possible in the identified areas.
Thanks to the deep pockets of governments and corporate giants, today’s research ecosystem is very good at supplying resources. However, when it comes to identifying impactful research, there are issues.
Today, it is university professors and professional scientists who direct and allocate these resources to promising areas of science. There are various mechanisms for this, such as grants which research proposals compete over or multi-year surveys where the academic community itself brainstorms the most promising research directions. Such mechanisms are well motivated. For example, by competing for grants, theoretically the proposals which the community thinks are the most promising should win.
However, all such mechanisms rely on one critical foundation — citations.
Citations are the currency of academia. When academics publish papers, they must cite the works they use. If a work is cited lots of times, it should be of higher value. If a scientist’s works are highly cited, that should be an indication that they are doing interesting and impactful science, and what they think matters. This gives them more social capital. As a result, the scientist gets more opportunities to review papers, grant proposals and also a more prominent role in community surveys. This way the most productive and capable researchers are in charge of shaping the future of their fields — at least theoretically. In the current system, citations are the key incentive as they give the primary indication for the “value” of a researcher.
Science’s Reproducibility Problem
The problem with this is that it misses the other crucial ingredient of good science — reproducibility.
Good research needs two things. Firstly, it should tackle interesting problems that people care about. This is where citations work. If many people cite a paper, it is clearly of interest to the community. Secondly, it must be reproducible. Other people must be able to reproduce your results with the methods presented in your papers. This is what confirms its validity and cements it as scientific fact.

Reproducibility is a fundamental and nonnegotiable part of the scientific method. If research is not independently verifiable, it is of no value no matter how stellar the results. If it cannot be reproduced, it is not science.
A central issue in today’s research ecosystem is that it overemphasises citations and largely just ignores reproducibility. Many claim that we are in the midst of a reproducibility crisis.
In many cases, this is not fatal. Scientists usually work in good faith and present honest results to the best of their ability. Papers are subject to peer review by multiple independent experts before being accepted by journals. These reviewers try their best to rigorously scrutinise submissions and probe them for inconsistencies. Finally, the works that cite a paper often cite it because their results would be physically impossible without the advances of the cited paper, which serves as a built-in check on reproducibility.
However, it would be naive to assume scientists universally operate in good faith. Examples of scientific misconduct and deception are abundant. Even if the underlying research is sound, there are examples of scientists forming “citation cartels” where they cite each other’s work to artificially amplify its value. Bad faith operators, while rare, compromise the value of the citation based system of identifying quality research/researchers. This is why citations alone are not enough, and reproducibility is essential.
For example, consider how much of the misinformation during the COVID-19 pandemic has masqueraded as “scientific fact”. Scientific papers put out in a rush with flawed methods — even with good intentions — have contributed to the misinformation crisis. As in broader academia, this is only true for a small minority of scientists studying COVID, but in a public health crisis even a small amount of bad science is dangerous. Given the intense attention on such research during the pandemic, it’s not hard for bad research to gain citations and appear reputable. If these bogus results instead had to be reproduced before being published, perhaps they would have been exposed sooner.
Make Reproducibility Easy Again
There is a problem though — reproducing results is hard. Many experiments require extremely specialised equipment that is expensive and difficult to get. For example, the discovery of the Higgs Boson at the Large Hadron Collider at CERN isn’t exactly independently reproducible unless someone is willing to fund another $5B particle accelerator. Even on smaller scales, quite often papers discuss the creation of entirely new devices with extremely specialised techniques. Having to reproduce such complicated work would create a huge drain on peer reviewers and prevent them from advancing their own research. The entire research ecosystem would slow to a snail’s pace, and the rapid innovation currently underway would be impossible. A blanket requirement to reproduce all research is clearly too extreme. We need to strike a balance.
A more sophisticated approach can vary the burden of reproducibility given the details of the individual papers. Let’s start with the category that should be the most easily reproducible — computational/mathematical research. What if in these papers, we could replace citations with import statements?
This idea, from Balaji Srinivasan (who has himself gone through the academic grind as a doctoral student at Stanford), is as follows. Computational research typically relies just on code and data. The paper is simply the human-readable form of the code and the data underlying it and it is the latter two parts that carry the novel contributions. There should therefore be a way to interact just with the code and data without needing to care about the paper and what journal/conference it is in. Essentially, we could treat such papers as pieces of software, and apply the principles of good software design to test for reproducibility.
One principle of software design is unit testing. Every single file has test cases that are used to make sure the code is doing what it says. By writing a robust array of tests that check all the main and edge cases, we can be fairly confident our code does what we intended for it to do without having to read it line by line. We could apply a similar technique to the code underlying papers by providing a set of tests to prove that the code produces the claimed results.
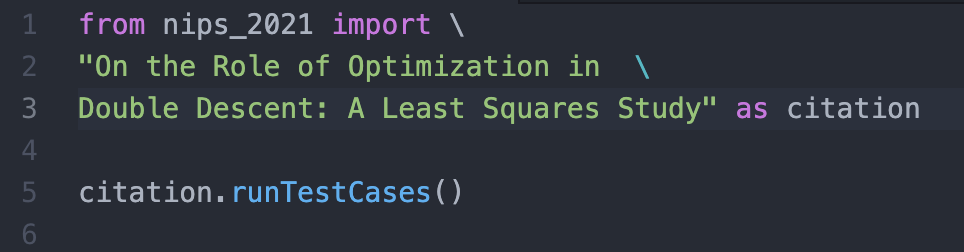
Suppose a researcher comes up with a new algorithm. Along with the code, they can supply a suite of test cases that illustrate that it works as desired. Then other researchers can simply “import” the unit tests into their own code to verify the work. With this, verifying reproducibility becomes easier. One loophole is if the tests supplied by the original author aren’t robust but this is offset by the fact that it’s easier to check that tests actually verify an algorithm rather than verify that an algorithm actually works. If the tests supplied aren’t meaningful, that automatically puts the prestige of the researcher on the line. In a system which values both reproducibility and citations, this provides a stabilising incentive for the author to provide robust test cases.
While this import-statement approach still has many details to be fleshed out, it isn’t so hard to imagine applications in computational and mathematical fields such as machine learning and statistics. One example of a detail that would need to be ironed out is where the code/data/test package would be stored. Maybe journals like Nature and Science might facilitate this transition themselves if they can get over their incumbent inertia. Srinivasan has also suggested independent blockchains1 to ensure a tamper proof record of research protected by cryptography.
For more mathematical proof based papers, computational ways to independently verify proofs already exist. One example is the open source software Lean, which is already being used in many applications. One example is the Xena Project at Imperial College London which is attempting to rewrite all the proofs from the undergraduate math curriculum using Lean. Independent computational verifiers like this offer a separate check on the logic of proofs beyond just the approval of highly cited researchers. This acts as a failsafe to protect against bad actors and ensure the integrity of mathematical research.
Reproducible Devices
It is much harder to imagine how we might extend this to research which is more than just code and instead involves new devices/machines. One interesting possibility, also laid out by Srinivasan, attempts to make it less prohibitive by once again taking inspiration from blockchains.
One of the data structures underlying a blockchain is a Merkle tree. Merkle trees can be visualised as tree structures where each node is composed of the cryptographic hash function of the nodes below it.
For experts, the TL;DR is that just like the hash at the top of the Merkle tree is the unique output of a unique combination of inputs at the bottom most layer, so too is the final device we produce the unique output of the unique way in which we join all the individual parts together. The Merkle tree therefore acts as a way to verify the specification the authors give in their paper and provides a basic verification without having to actually make the device from scratch.
For the rest of us, here is a breakdown of what a Merkle tree is and how it actually helps us with this reproducibility problem.
A hash function is a function that takes an input of arbitrary size to a fixed size output, e.g. it could take any word and map it to a number. If f is our function, then maybe f(“apple”) = 1, f(“banana”) = 2 etc. The challenge is to come up with such a function so that each unique input gives a unique output, but at the same time you can’t easily get the input from the output value.
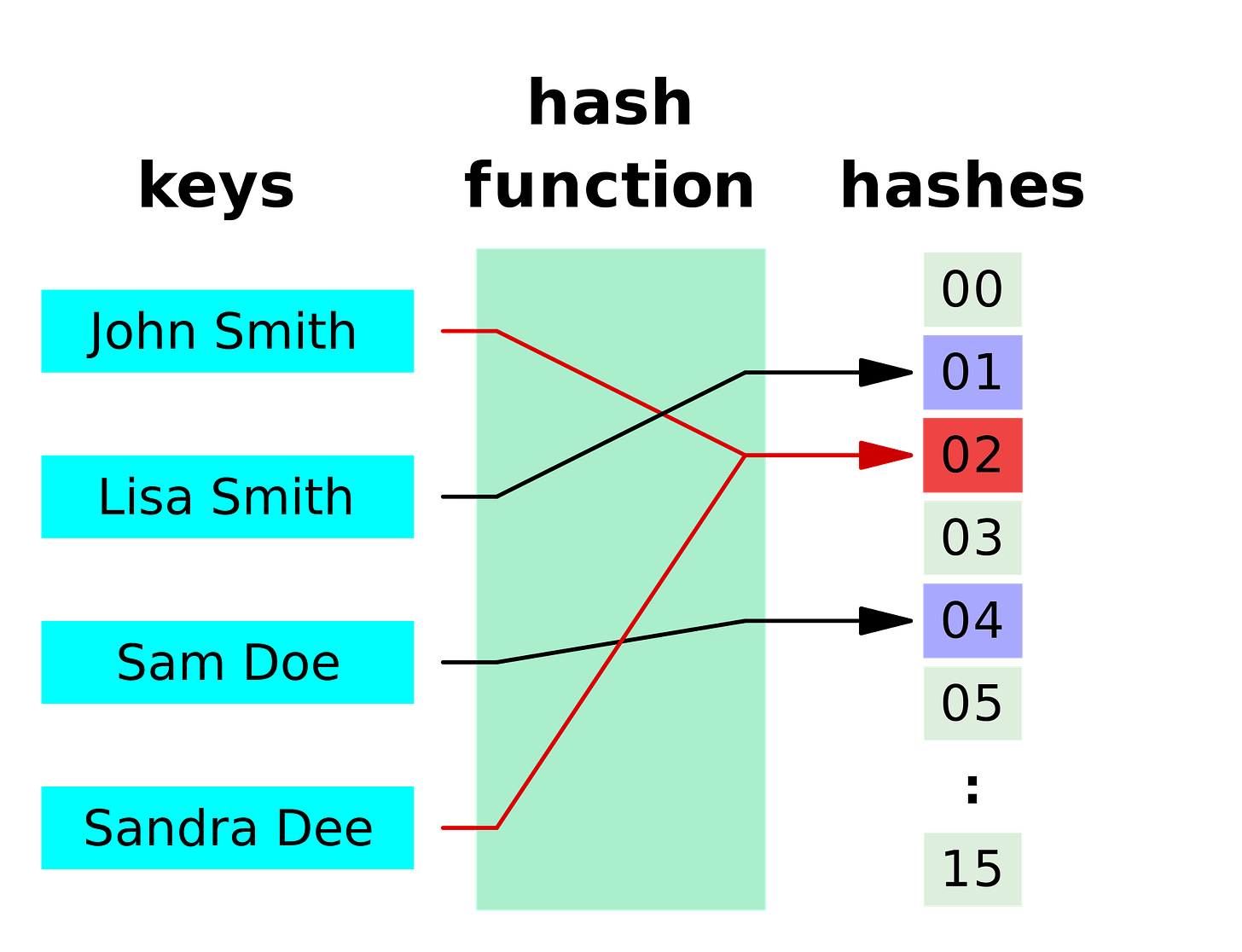

For example, if our hash function assigned each word to its length, then f(“apple”) = 5 and f(“banana”) = 6. However, this wouldn’t be a great function since plenty of words would have the same output, e.g. f(“pear”) = f(“fear”) = f(“park”) = 4. Fortunately, there are plenty of good readymade hash functions which we can use.
The next component is a tree. This is a simple structure for storing data which has a (surprise surprise) tree-like hierarchy. There is a node at the top of the tree, which has its own children nodes below it, and the children nodes have nodes below them, until we come to the bottom most layer.
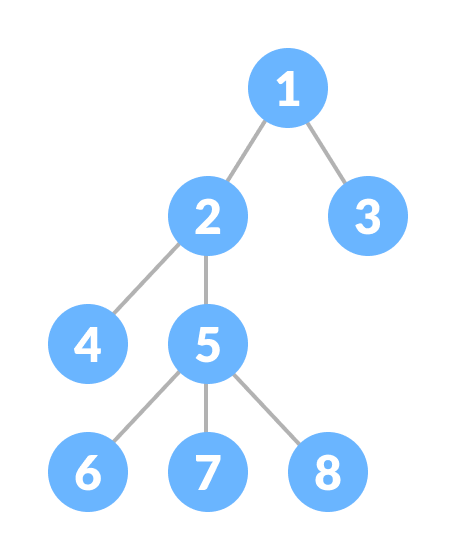
In a Merkle tree, the data in the each of the nodes isn’t just random numbers, but the hash function of the data in its children nodes. The nodes at the very bottom have a preset value and correspond to the inputs of the Merkle tree. By combining the nodes at the bottom in different ways, we end up with a unique hash value at the top most node.
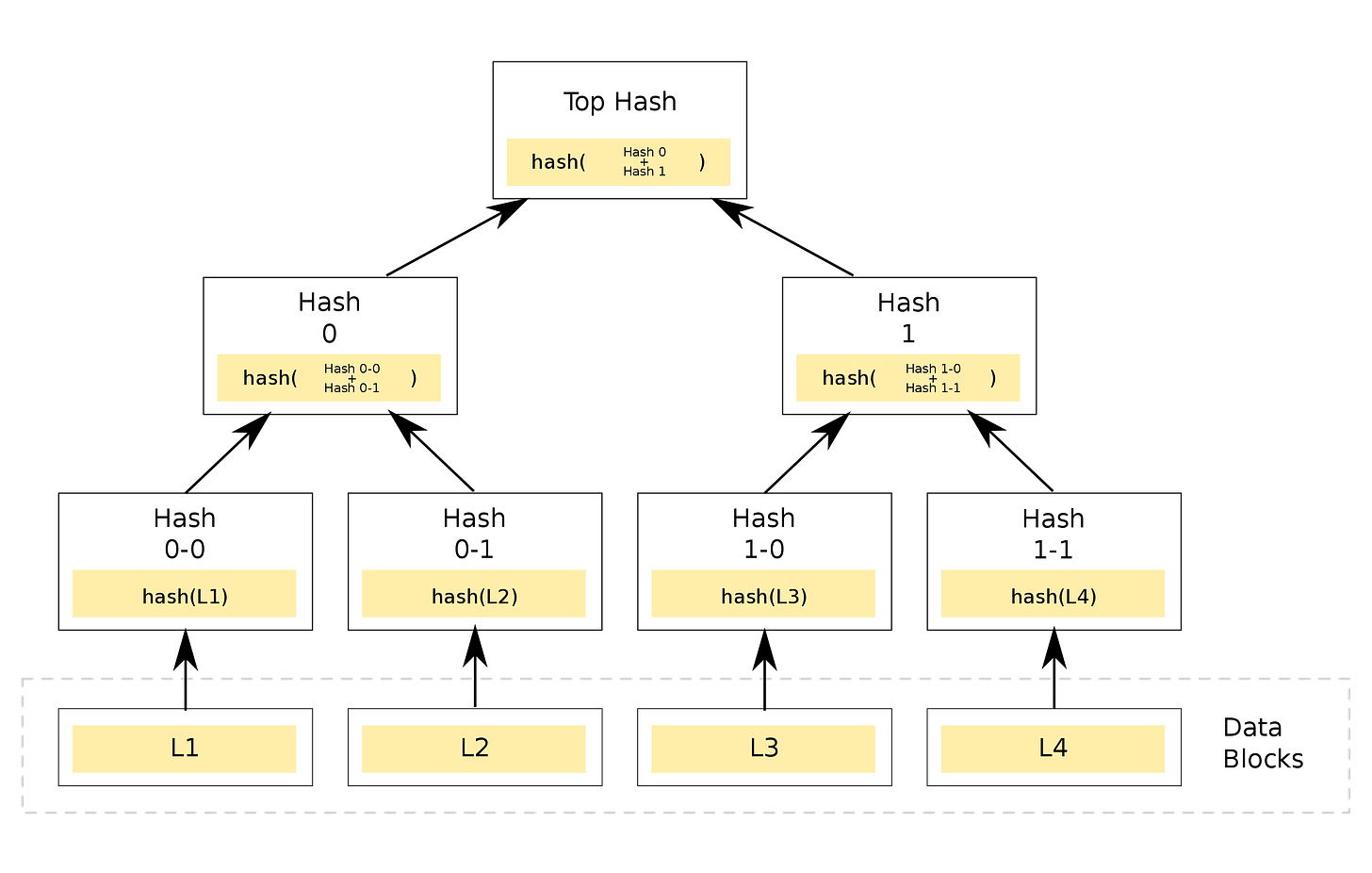
With this, we can come up with a basic idea for verifiably transmitting instructions to make new devices. For a toy example to illustrate the idea, suppose we are making a basic device which has a lightbulb and a battery connected by a copper wire. First, let’s hash the words “lightbulb”, “battery”, and “copper wire” using the hash function SHA-256, which outputs a 64 digit hexadecimal number:
lighbulb: beed9cd9613ee690c73dc1cc968064d8387cd42385694a4ddb22d09615921ab0
battery: f3d1701e1d575e1294786989517866986bc97343e07af63e201f46ba0be5806a
copper wire: 60cc0539bc9b6538b027b592002addcdb8796e298e29f52a079206fa15193417
Suppose we first combine the battery and the wire, and then add the lightbulb. For the combination step, we can choose to take the hash of the two underlying strings concatenated. For example combining the battery and wire means we need to compute SHA-256 for the input “beed9cd9613ee690c73dc1cc968064d8387cd42385694a4ddb22d09615921ab0f3d1701e1d575e1294786989517866986bc97343e07af63e201f46ba0be5806a”.
Doing this gives the following Merkle tree:
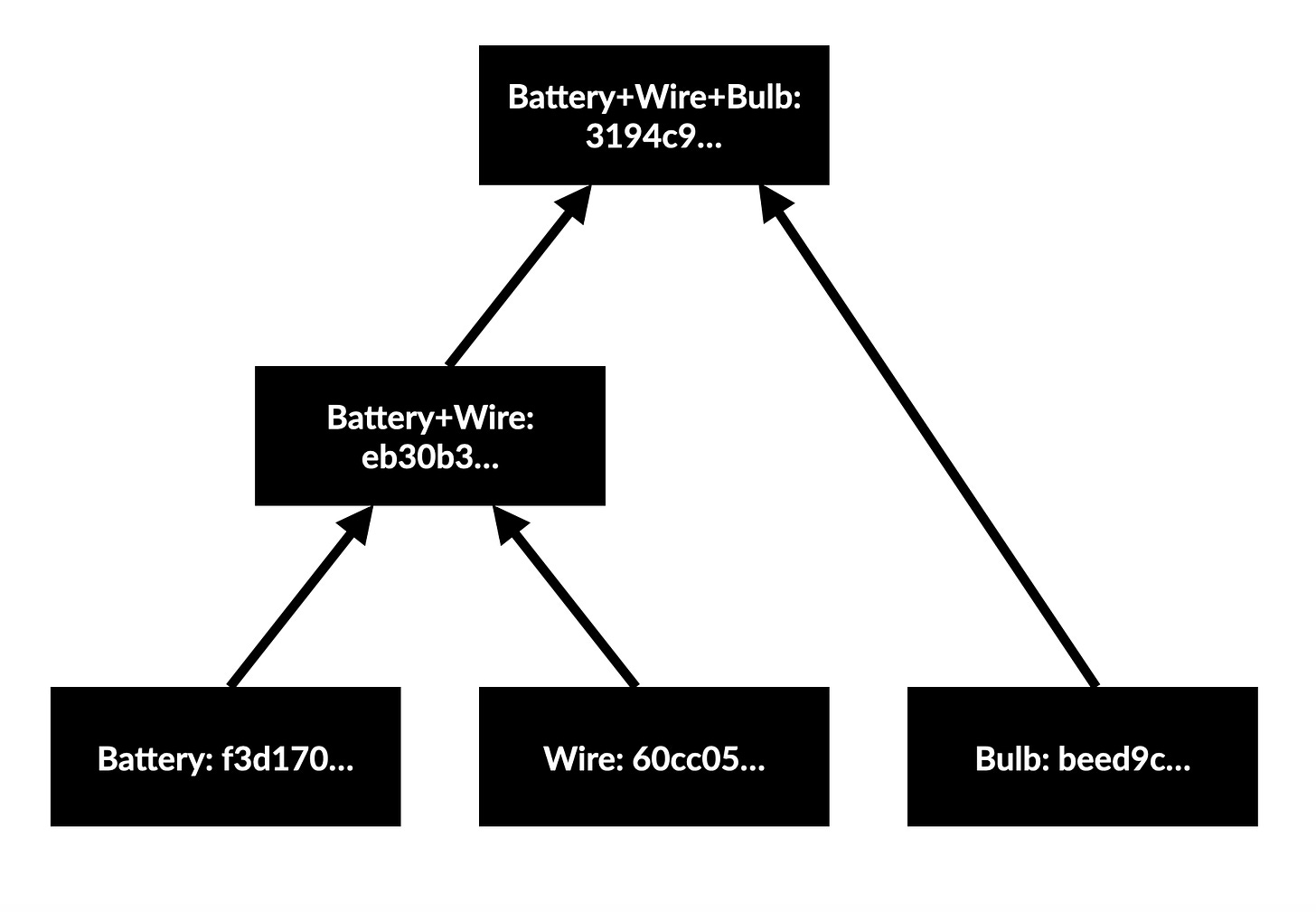
Our final circuit has the unique hash value beginning 3194c9… which could only be produced if we combine the components with those initial hashes in that exact order. If we had instead joined wire and bulb first then added the battery, the final hash would be c6a8c4… Of course in this case the order which we join parts doesn’t really matter so both of these should give the same hash, but that detail can be sorted by choosing a more sophisticated hashing function/intelligently assigning input hashes.
One wrinkle to experiment with is the way in which hashes are combined. Our naive implementation assumed there was only one way to join components, represented by only one way to combine their hashes but we could have different hash combinations depending on which methods we use to join parts.
The key is that by identifying our starting components with numbers, we can use a Merkle tree to track them and produce the unique output hash of the device. Then another researcher can be convinced that the procedure set out in the paper is correct as that is the only one that can produce the final device hash given all the input part hashes. The author’s specification is verified without actually having to physically create the new device, as long as the author and the reviewer agree on exactly what components the initial hashes refer to, which can be done by creating a common database of components. This allows us to sidestep the difficulty of actually reproducing the device and the large time cost cost associated with that.
This is far from the ideal of actually building the device and manually verifying results, but it offers some degree of accountability without totally paralysing research. Industrial companies already assign serial numbers to uniquely identify their components, and academia chould consider experimenting with the idea also.
Nevertheless, there are lots of difficulties that might make this very difficult. It would require systematic categorisation of the vast array of components/fabrication techniques that researchers use. There is also the possibility that a malicious author gives a specification in their paper that passes the Merkle tree test, but then actually uses another device entirely to produce their results. In such cases, Merkle trees will be useless without an actual attempt at replication. Nevertheless, even if this idea is too simplistic it is at least an interesting proposal to build on to make independent verification more feasible for non computational research.
There are two aspects of quality research. The first is importance and this is parametrised well by citations. The second is reproducibility, which cements research as verified. However, today the academic community has become too reliant on citations as an indicator and ignored reproducibility. In most cases where scientists work in good faith, this is passable. However, bad faith actors have used this to inflate the importance of their own research through tricks like citation cartels. At its worst, the race for citations has contributed to misinformation via hastily designed experiments seeking to capitalise on a once in a generation pandemic.
The traditional reason for ignoring reproducibility is that it is very resource and time intensive. There is a tradeoff between reproducibility and progress, and till now, we have tilted far towards progress and dangerously ignored reproducibility. However, this is where new proposals such as Srinivasan’s software-like papers and device Merkle trees provide interesting alternatives. By leveraging such techniques, we could position ourselves more favourably on the reproducibility-progress tradeoff. Instead of completely neglecting it outright, we give reproducibility the respect it deserves while not sacrificing the pace of progress. Maybe these ideas are too simplistic, but clearly something is needed. It won’t be easy, but in the long run ignoring reproducibility is a very dangerous idea for science.
For those unfamiliar, blockchains are an absolutely fascinating innovation comparable to a shared database. The unique feature of blockchains is the requirement of consensus — even if A and B don’t trust each other, they will necessarily agree on the data stored in the blockchain they share due to the mathematical structure behind it.




